 |
UNIQORN -- The Universal Neural-network Interface for Quantum Observable Readout from N-body wavefunctions
|
 |
UNIQORN -- The Universal Neural-network Interface for Quantum Observable Readout from N-body wavefunctions
|
Functions | |
| def | data_ID_loader (**kwargs) |
| IMPORT DATASET AND LABELS. More... | |
| def | label_ID_loader (DataNameComplete, **kwargs) |
| def | extractor (WhichData, DataFileName, K) |
| def | DataThresholdSelection (IDs_tot, Low, High, NDatasets) |
Variables | |
| content = f.readline() | |
| extract single shots More... | |
| no_of_columns = len(content.split(' ')) | |
| NShots | |
| ThisData = np.transpose(np.loadtxt(DataFileName, usecols=np.arange(3,no_of_columns))) | |
| extract density More... | |
| Ntot = DataFileName.split('00N')[1].split('M')[0] | |
| extract particle number More... | |
| lines = f.read().splitlines() | |
| extract fragmentation More... | |
| last_line = lines[-1] | |
| FRAG = last_line.split(' ')[1].strip() | |
| _ | |
| extract phase histograms More... | |
| int | toskip = 2 |
| Xmin = str(min(np.loadtxt(DataFileName, usecols=[0],skiprows=toskip))) | |
| Xmax = str(min(np.loadtxt(DataFileName, usecols=[0],skiprows=toskip))) | |
| file_x_boundaries = open('Xboundaries.txt', 'w+') | |
| VizGrid = np.loadtxt(DataFileName, usecols=[0],skiprows=toskip) | |
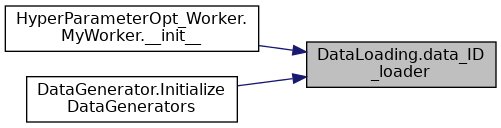
| def DataLoading.data_ID_loader | ( | ** | kwargs | ) |
IMPORT DATASET AND LABELS.
Loads the paths (IDs) for the dataset and already splits them into training and test sets.
Parameters
----------
**kwargs :
----------
- data (=inp.Training_Data) : str
specifies what is the data, can be any of the following:
* FRAG :
Flag to extract the fragmentation
* NPAR :
Flag to extract the number of particles
* DENS:
Flag to extract the density
* CORR1/CORR2/RHO1/RHO2:
Flag to extract the one-body or two-body reduced density matrix (RHO) or Galuber correlations (CORR)
* POT:
Flag to extract the potential
* PHASEHIST:
Flag to extract the phase histogram
Returns
-------
(IDs_training, IDs_test)
The paths where the data sets for training and test are stored.
References
----------
See Also
--------
Notes
-----
Examples
-------- 
| def DataLoading.DataThresholdSelection | ( | IDs_tot, | |
| Low, | |||
| High, | |||
| NDatasets | |||
| ) |


| def DataLoading.extractor | ( | WhichData, | |
| DataFileName, | |||
| K | |||
| ) |
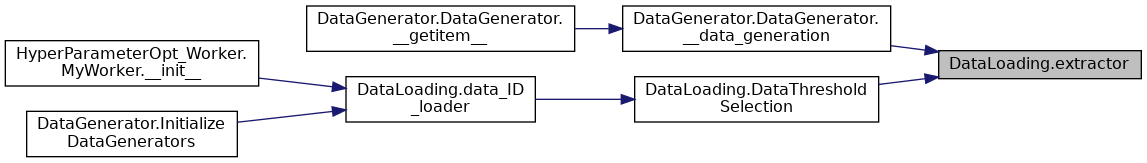
| def DataLoading.label_ID_loader | ( | DataNameComplete, | |
| ** | kwargs | ||
| ) |
Loads the paths (IDs) for the dataset and already splits them into training and test sets.
Parameters
----------
**kwargs :
----------
- labels (=inp.Learn) : str
specifies what is the data, can be any of the following:
* FRAG :
Flag to extract the fragmentation
* NPAR :
Flag to extract the number of particles
* DENS:
Flag to extract the density
* CORR1/CORR2/RHO1/RHO2:
Flag to extract the one-body or two-body reduced density matrix (RHO) or Galuber correlations (CORR)
* POT:
Flag to extract the potential
* PHASEHIST:
Flag to extract the phase histogram
Returns
-------
LabelNameComplete:
The paths where the labels of the corresponding data sets (for training or testing) are stored.
References
----------
See Also
--------
Notes
-----
Examples
-------- 
|
private |
extract phase histograms
| DataLoading.content = f.readline() |
extract single shots
| DataLoading.file_x_boundaries = open('Xboundaries.txt', 'w+') |
| string DataLoading.FRAG = last_line.split(' ')[1].strip() |
| DataLoading.last_line = lines[-1] |
| DataLoading.lines = f.read().splitlines() |
extract fragmentation
| DataLoading.no_of_columns = len(content.split(' ')) |
| DataLoading.NShots |
| DataLoading.Ntot = DataFileName.split('00N')[1].split('M')[0] |
extract particle number
| tuple DataLoading.ThisData = np.transpose(np.loadtxt(DataFileName, usecols=np.arange(3,no_of_columns))) |
extract density
extract potential
extract correlations
calculate/extract the correlation functions or reduced density matrix form MCTDHX data
| int DataLoading.toskip = 2 |
| DataLoading.VizGrid = np.loadtxt(DataFileName, usecols=[0],skiprows=toskip) |
| DataLoading.Xmax = str(min(np.loadtxt(DataFileName, usecols=[0],skiprows=toskip))) |
| DataLoading.Xmin = str(min(np.loadtxt(DataFileName, usecols=[0],skiprows=toskip))) |